|
See the GNU General Public License for more details. This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version. Export trace or sequence to different format.Open the "Find Sequence." from the edit menu Search for a regular expression pattern.Display Sequence and metadata from the view menu.Make the reverse complement from the edit menu.Then you can cut or copy to the clipboard Select a subsequence with the mouse as with any text editor.Scale the trace horizontally or vertically using 2 sliders at the bottom right.( Finger geasture are supported with touch screen) Once the file is open, cutepeaks allows you to : You can open those files from cutepeaks by clicking on open from the File menu. Installation Windowsįor ubuntu >=21.04, Download this one hereĬutePeaks supports following trace file formats: However, it is written with GTK framework in Python 2, the latter being deprecated and slower than C++. Seqtrace is the only standalone and opensource application we could find. 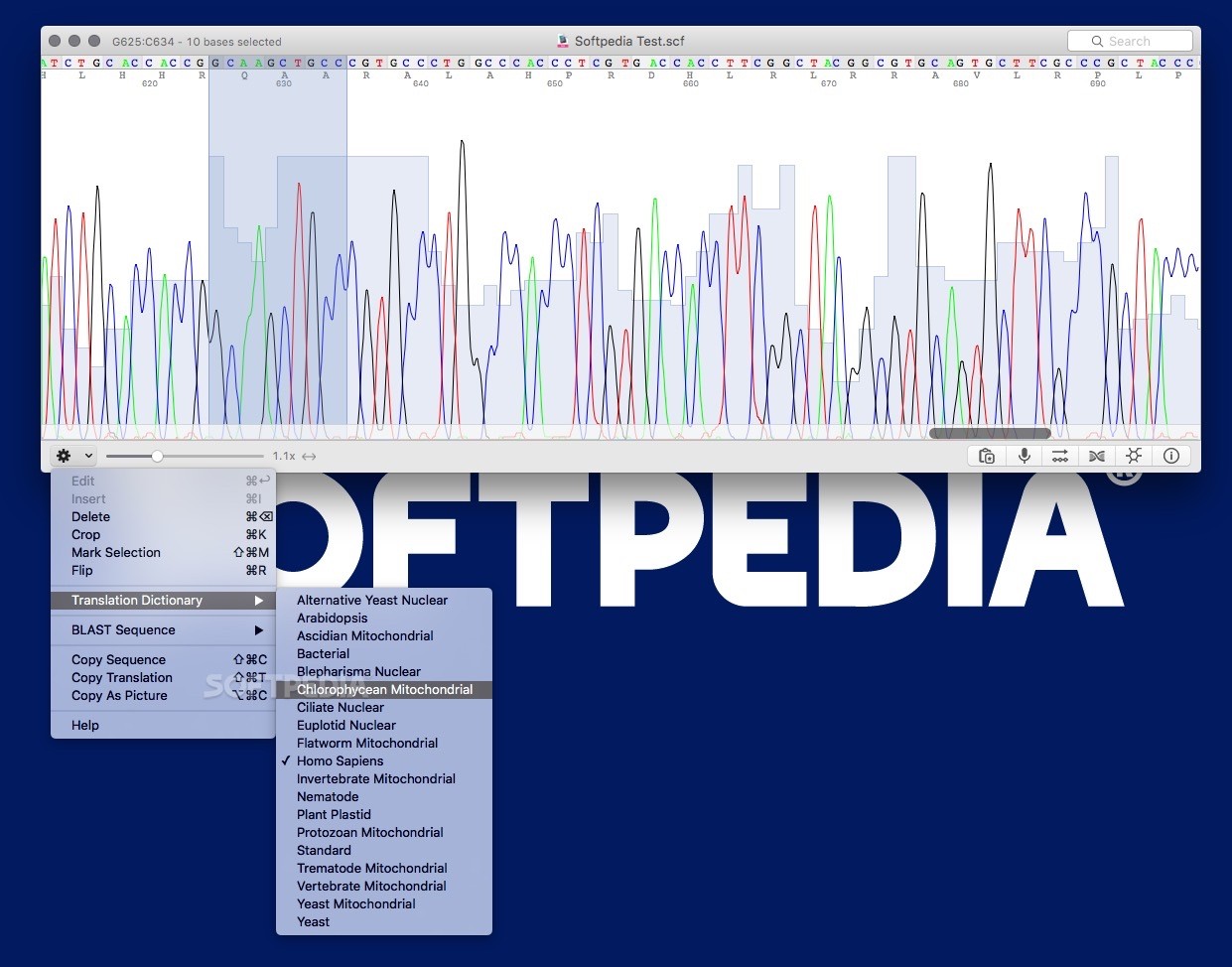 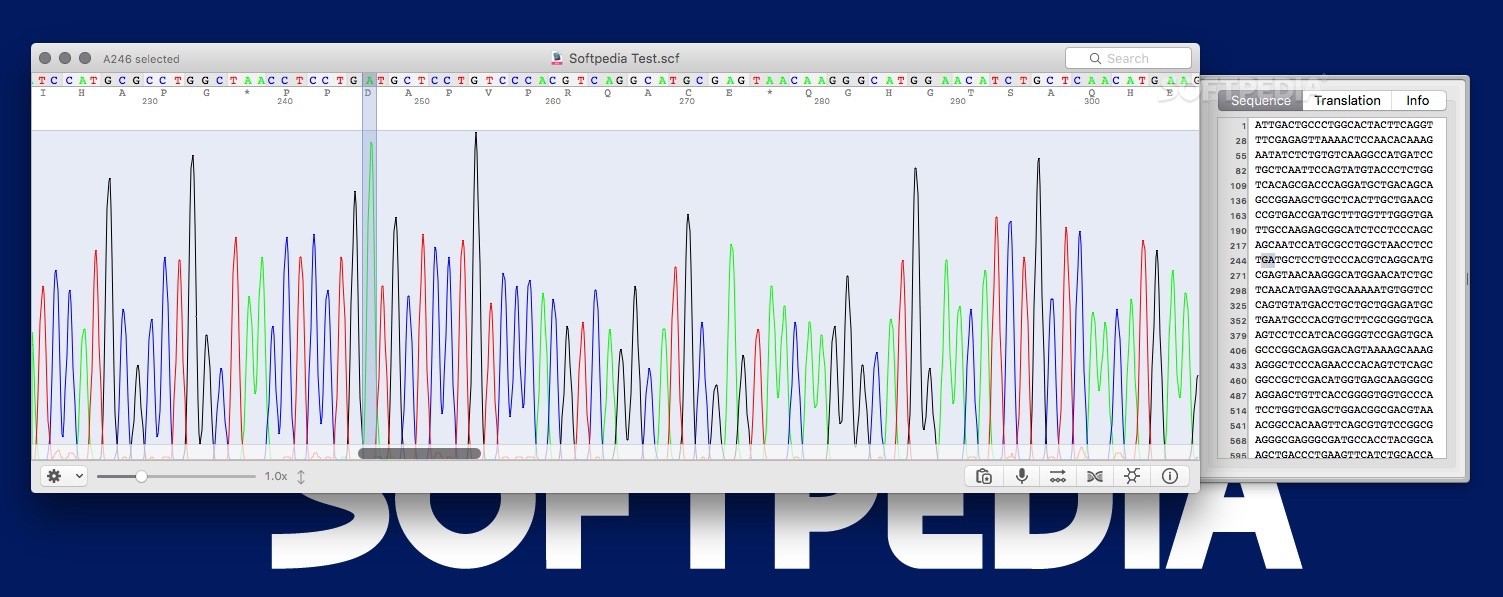
Sadly, it is only available on MacOS and source code is not opened to community enhancement. State of fieldĤpeaks is a software widely used by biologists that benefits from a nice User interface. Moreover, they are not always user-friendly and lack modern look and feel. 
Very few opensource software is available to explore Sanger trace data and most of labs staff still rely on proprietary software. It has regular expression pattern finder and can export trace as SVG vector image.ĭespite the major use of Next Generation Sequencing, the Sanger method is still widely used in genetic labs as the gold standard to read target DNA sequences. A simple viewer for Sanger trace file made with Qt5.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
